This document is relevant for: Trn2, Trn3
Profile a NKI Kernel#
Learn how to profile Neuron Kernel Interface (NKI) kernels using Neuron Explorer to analyze hardware-level performance characteristics on Trainium and Inferentia devices. This comprehensive guide covers two profiling methods: using the neuron-explorer capture command-line tool. You’ll discover how to generate NEFF and NTFF files, identify performance bottlenecks, optimize kernel execution, and leverage the interactive web-based Neuron Profile UI to visualize execution traces with source code integration for efficient NKI kernel development and optimization.
Install Neuron Explorer#
Ensure that you have the latest version of the aws-neuronx-tools package installed as Neuron Explorer comes with this package. The aws-neuronx-tools package is pre-installed on Neuron DLAMIs.
For detailed installation instructions, see: How to Get Started with Neuron Explorer.
Profile a NKI Kernel#
Profiling NKI (Neuron Kernel Interface) kernels helps you understand hardware level performance characteristics of your kernels running on AWS Trainium and Inferentia devices. When you write or optimize custom NKI kernels, profiling allows you to:
Identify bottlenecks: Determine if your kernel is compute-bound, memory-bound, or limited by data movement.
Optimize performance: Analyze kernel-level execution time, investigate compute engine utilization, look for opportunities to implement operator fusion to fine-tune performance.
Compare implementations: Benchmark different kernel implementations or configurations to pick the most efficient kernel.
You can profile NKI kernels using several approaches. In this guide, you’ll learn two primary methods for profiling NKI kernels.
How to profile using neuron-explorer capture#
To profile an NKI kernel using neuron-explorer capture, follow these three steps:
Set the environment variable
NEURON_FRAMEWORK_DEBUG=1to instruct the compiler to save the NEFF (Neuron Executable File Format) file.Execute the NKI kernel to generate the NEFF file.
Run
neuron-explorer captureto create an Neuron Trace File Format (NTFF) file for performance analysis.
Each of these steps is explained in detail below.
Step 1: Set Environment Variables#
We will profile a 3-layer MLP model that fuses matrix multiplications with ReLU activation functions and uses a NKI matrix multiplication kernel. The rest of this tutorial will use a performance profile generated from this example. Here is the implementation of mlp_with_mm_kernel.py. Save this file before moving on to the next step:
"""
Example 3-layer MLP with matrix multiplication kernel to demonstrate Neuron Profile.
"""
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch_neuronx
import nki
import nki.isa as nisa
import nki.language as nl
import os
os.environ["NEURON_LOGICAL_NC_CONFIG"] = "1"
os.environ["NEURON_FRAMEWORK_DEBUG"] = "1"
os.environ["XLA_IR_DEBUG"] = "1" # Preserve source-level IR names in the compiled graph for profiler source mapping
os.environ["XLA_HLO_DEBUG"] = "1" # Preserve HLO operation names and metadata for profiler attribution
@nki.jit
def nki_matmul(
lhsT,
rhs,
# Meta-parameters
TILES_IN_BLOCK_M=16,
TILES_IN_BLOCK_N=2,
TILES_IN_BLOCK_K=8,
):
"""NKI kernel to compute a large matrix multiplication efficiently by
blocking all dimensions and doing layout optimization.
Args:
lhsT: an input tensor of shape [K,M], where K is a multiple of 128 *
TILES_IN_BLOCK_K and M is a multiple of 128 * TILES_IN_BLOCK_M. It is the
left-hand-side argument of the matrix multiplication, delivered transposed
for optimal performance.
rhs: an input tensor of shape [K,N], where K is a multiple of 128 *
TILES_IN_BLOCK_K and N is a multiple of 512 * TILES_IN_BLOCK_N. It is
the right-hand-side argument of the matrix multiplication.
TILES_IN_BLOCK_*: meta parameters to control blocking dimensions
Returns:
result: the resulting output tensor of shape [M,N]
"""
# Verify that the lhsT and rhs have the same contraction dimension.
K, M = lhsT.shape
K_, N = rhs.shape
assert K == K_, "lhsT and rhs must have the same contraction dimension"
# Lookup the device matrix multiply dimensions.
TILE_M = nl.tile_size.gemm_stationary_fmax # 128
TILE_K = nl.tile_size.pmax # 128
TILE_N = nl.tile_size.gemm_moving_fmax # 512
# Compute the block dimensions.
BLOCK_M = TILE_M * TILES_IN_BLOCK_M
BLOCK_N = TILE_N * TILES_IN_BLOCK_N
BLOCK_K = TILE_K * TILES_IN_BLOCK_K
# Verify the size is a multiple of block size
assert M % BLOCK_M == 0, \
f"Expected M {M} to be divisible by {BLOCK_M} when there are {TILES_IN_BLOCK_M}"
assert N % BLOCK_N == 0, \
f"Expected N {N} to be divisible by {BLOCK_N} when there are {TILES_IN_BLOCK_N}"
assert K % BLOCK_K == 0, \
f"Expected K {K} to be divisible by {BLOCK_K} when there are {TILES_IN_BLOCK_K}"
# Create a space for the result in HBM (not initialized)
result = nl.ndarray((M, N), dtype=lhsT.dtype, buffer=nl.shared_hbm)
# Compute the number of blocks in each dimension
NUM_BLOCK_M = M // BLOCK_M
NUM_BLOCK_N = N // BLOCK_N
NUM_BLOCK_K = K // BLOCK_K
# Blocking N dimension (the RHS free dimension)
for n in nl.affine_range(NUM_BLOCK_N):
n_start = n * BLOCK_N
n_end = n_start + BLOCK_N
# Allocate and initialize result matrix N-block to 0.0.
#
# Each result M-tile stores its N-block contiguous on the free-dim
# with shape (TILE_M, TILES_IN_BLOCK_N, TILE_N). This layout allows
# reshaping to (TILE_M, BLOCK_N) for SBUF->HBM DMA to operate on a
# large payload, enabling good DMA efficiency.
#
# We split the N-block into individual M-tiles so the compiler can
# pipeline memset(0), matmul, tensor_tensor, and SBUF->HBM DMA
# on M-tile granularity.
result_m_tiles = []
for m in nl.affine_range(NUM_BLOCK_M):
for m_tile in nl.affine_range(TILES_IN_BLOCK_M):
result_m_tile = nl.ndarray(
shape=(TILE_M, TILES_IN_BLOCK_N, TILE_N),
dtype=result.dtype,
buffer=nl.sbuf,
)
nisa.memset(dst=result_m_tile, value=0.0)
result_m_tiles.append(result_m_tile)
# Blocking K dimension (the contraction dimension)
for k in nl.sequential_range(NUM_BLOCK_K):
k_block_tile_start = k * TILES_IN_BLOCK_K
# Load tiles from RHS
# Load tiles one N-block at a time for good DMA efficiency.
rhs_tiles = nl.ndarray(
shape=(TILE_K, TILES_IN_BLOCK_K, BLOCK_N),
dtype=rhs.dtype,
buffer=nl.sbuf,
)
for k_tile in range(TILES_IN_BLOCK_K):
k_tile_start = (k_block_tile_start + k_tile) * TILE_K
k_tile_end = k_tile_start + TILE_K
nisa.dma_copy(
dst=rhs_tiles[0:TILE_K, k_tile, 0:BLOCK_N],
src=rhs[k_tile_start:k_tile_end, n_start:n_end],
)
# Blocking M dimension (the LHS free dimension)
for m in nl.affine_range(NUM_BLOCK_M):
# Loading tiles from lhsT
# Load tiles one M-block at a time for good DMA efficiency.
lhsT_tiles = nl.ndarray(
shape=(TILE_K, TILES_IN_BLOCK_K, BLOCK_M),
dtype=lhsT.dtype,
buffer=nl.sbuf,
)
m_start = m * BLOCK_M
m_end = m_start + BLOCK_M
for k_tile in nl.affine_range(TILES_IN_BLOCK_K):
k_tile_start = (k_block_tile_start + k_tile) * TILE_K
k_tile_end = k_tile_start + TILE_K
nisa.dma_copy(
dst=lhsT_tiles[0:TILE_K, k_tile, 0:BLOCK_M],
src=lhsT[k_tile_start:k_tile_end, m_start:m_end],
)
# Do matmul with all tiles in the blocks
m_block_tile_start = m * TILES_IN_BLOCK_M
for n_tile in nl.affine_range(TILES_IN_BLOCK_N):
for m_tile in nl.affine_range(TILES_IN_BLOCK_M):
result_tile = nl.ndarray(
shape=(TILE_M, TILE_N), dtype=nl.float32, buffer=nl.psum
)
for k_tile in nl.affine_range(TILES_IN_BLOCK_K):
m_tile_start = m_tile * TILE_M
m_tile_end = m_tile_start + TILE_M
n_tile_start = n_tile * TILE_N
n_tile_end = n_tile_start + TILE_N
nisa.nc_matmul(
dst=result_tile,
stationary=lhsT_tiles[0:TILE_K, k_tile, m_tile_start:m_tile_end],
moving=rhs_tiles[0:TILE_K, k_tile, n_tile_start:n_tile_end],
)
# Evict from PSUM to SBUF while accumulating into result M-tile.
m_tile_idx = m_block_tile_start + m_tile
result_m_tile = result_m_tiles[m_tile_idx]
nisa.tensor_tensor(
dst=result_m_tile[0:TILE_M, n_tile, 0:TILE_N],
data1=result_m_tile[0:TILE_M, n_tile, 0:TILE_N],
data2=result_tile,
op=nl.add,
)
# Evict the result M-tiles from SBUF to HBM.
# Copy on N-blocks granularity for good DMA efficiency.
for m in nl.affine_range(NUM_BLOCK_M):
m_block_tile_start = m * TILES_IN_BLOCK_M
for m_tile in nl.affine_range(TILES_IN_BLOCK_M):
m_tile_idx = m_block_tile_start + m_tile
result_m_tile = result_m_tiles[m_tile_idx]
result_m_tile_block = result_m_tile.reshape((TILE_M, BLOCK_N))
m_tile_start = m_tile_idx * TILE_M
m_tile_end = m_tile_start + TILE_M
nisa.dma_copy(
dst=result[m_tile_start:m_tile_end, n_start:n_end],
src=result_m_tile_block[0:TILE_M, 0:BLOCK_N],
)
return result
class NKILinear(nn.Module):
def __init__(self, in_features, out_features):
super(NKILinear, self).__init__()
self.weight = nn.Parameter(torch.randn(out_features, in_features))
self.bias = nn.Parameter(torch.randn(out_features))
def forward(self, x):
weight_T = self.weight.t()
x_T = x.t()
output = nki_matmul(x_T, weight_T)
return output + self.bias
class MLP(nn.Module):
def __init__(self):
super(MLP, self).__init__()
self.fc1 = NKILinear(2048, 2048)
self.fc2 = NKILinear(2048, 1024)
self.fc3 = NKILinear(1024, 1024)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return F.log_softmax(x, dim=1)
def main():
torch.manual_seed(0)
model = MLP()
train_x = torch.randn(2048, 2048)
# Use torch_neuronx.trace to compile the model and generate the NEFF
traced_model = torch_neuronx.trace(model, train_x, compiler_args="--lnc=1", compiler_workdir="./compiler_workdir")
output = traced_model(train_x)
print(f"Output tensor: {output}")
if __name__ == "__main__":
main()
As you can see, at the very top we have added the following environment variables:
os.environ["NEURON_FRAMEWORK_DEBUG"] = "1"
os.environ["XLA_IR_DEBUG"] = "1"
os.environ["XLA_HLO_DEBUG"] = "1"
These environment variables serve the following purposes:
NEURON_FRAMEWORK_DEBUG=1: Enables Neuron debug output. This triggers the Neuron compiler to save the Neuron Executable File Format (NEFF) artifact to the current directory after compilation of your NKI kernel. The NEFF contains all hardware instructions required to execute your NKI kernel on a NeuronDevice, as well as metadata and debug info needed for profiling.XLA_IR_DEBUG=1: Preserves the mapping between high-level framework operations (e.g., PyTorch operators) and the intermediate representation (IR) passed to the compiler. This enables source code linking from device instructions back to framework-level code in the profiler.XLA_HLO_DEBUG=1: Preserves the mapping between the HLO (High Level Operation) graph and the original framework operations. This enables the profiler to display descriptive operator names and stack frame information, making it easier to identify which part of your model corresponds to each device instruction.
Step 2: Compile Your NKI Kernel#
Compile your NKI kernel to create a NEFF in your current directory:
$ python3 mlp_with_mm_kernel.py
Note
The compiler_workdir argument to torch_neuronx.trace specifies the directory where the compiler saves artifacts, including the NEFF file. Look for your NEFF file inside the ./compiler_workdir directory, which will be named graph.neff.
Step 3: Profile the Generated NEFF#
The last step is profiling the generated NEFF. This step executes the NEFF on the NeuronDevice and records a raw execution trace into a NTFF artifact:
$ neuron-explorer capture -n ./compiler_workdir/graph.neff -s profile.ntff --profile-nth-exec=2 --enable-dge-notifs
This will save your NTFF profile to profile_exec_2.ntff.
Important
The --profile-nth-exec=2 option will profile your NEFF twice on the NeuronDevice and output a NTFF profile for the second iteration. This is recommended to avoid one-time warmup delays which can be seen in the first iteration of execution.
The --enable-dge-notifs option enables the capture of DGE DMA events but has known issues where it may overflow the status notification queue and cause execution timeouts when there are many DGE instructions.
View the Neuron Explorer UI#
This section assumes you’ve completed the previous step and have already generated both the NEFF and NTFF files, and downloaded them on your local machine.
Neuron Explorer includes an interactive, web-based UI for exploring execution traces in detail. In this section, we’ll open the Neuron Explorer UI to examine NKI-specific profiling information. These details can be found in multiple areas of the interface — including instruction hover tooltips, instruction click panels, search results, and box select results. For a comprehensive overview of all available viewers, see the Neuron Explorer documentation.
To view the Neuron Profile Web UI, execute the view command to start Web UI, replacing <workspace> with a path to a folder to store your profiling artifacts:
$ neuron-explorer view --data-path ./<workspace>
<workspace> is a path that neuron profile will use for storing and managing profiles.
The above command should print a URL that you can click to open the web UI:
View a list of profiles at http://localhost:3001/
Port Forwarding for Remote Instances#
If neuron-explorer view is run on a remote instance, you may need to use port forwarding to access the web UI. By default, neuron-explorer creates a web server on port 3001 and the API server on port 3002. To enable connection to your browser in you local computer, we will need to establish an ssh tunnel to both of the ports.
For example:
ssh -L 3001:localhost:3001 -L 3002:localhost:3002 <user>@<ip> -fN
If you created an EC2 instance with pem credentials, include it in the ssh tunnel below:
ssh -i ~/my-ec2.pem -L 3001:localhost:3001 -L 3002:localhost:3002 ubuntu@[PUBLIC_IP_ADDRESS] -fN
Using the Profile UI#
Once the ssh tunnel is setup, you can now open a browser and navigate to http://localhost:3001.
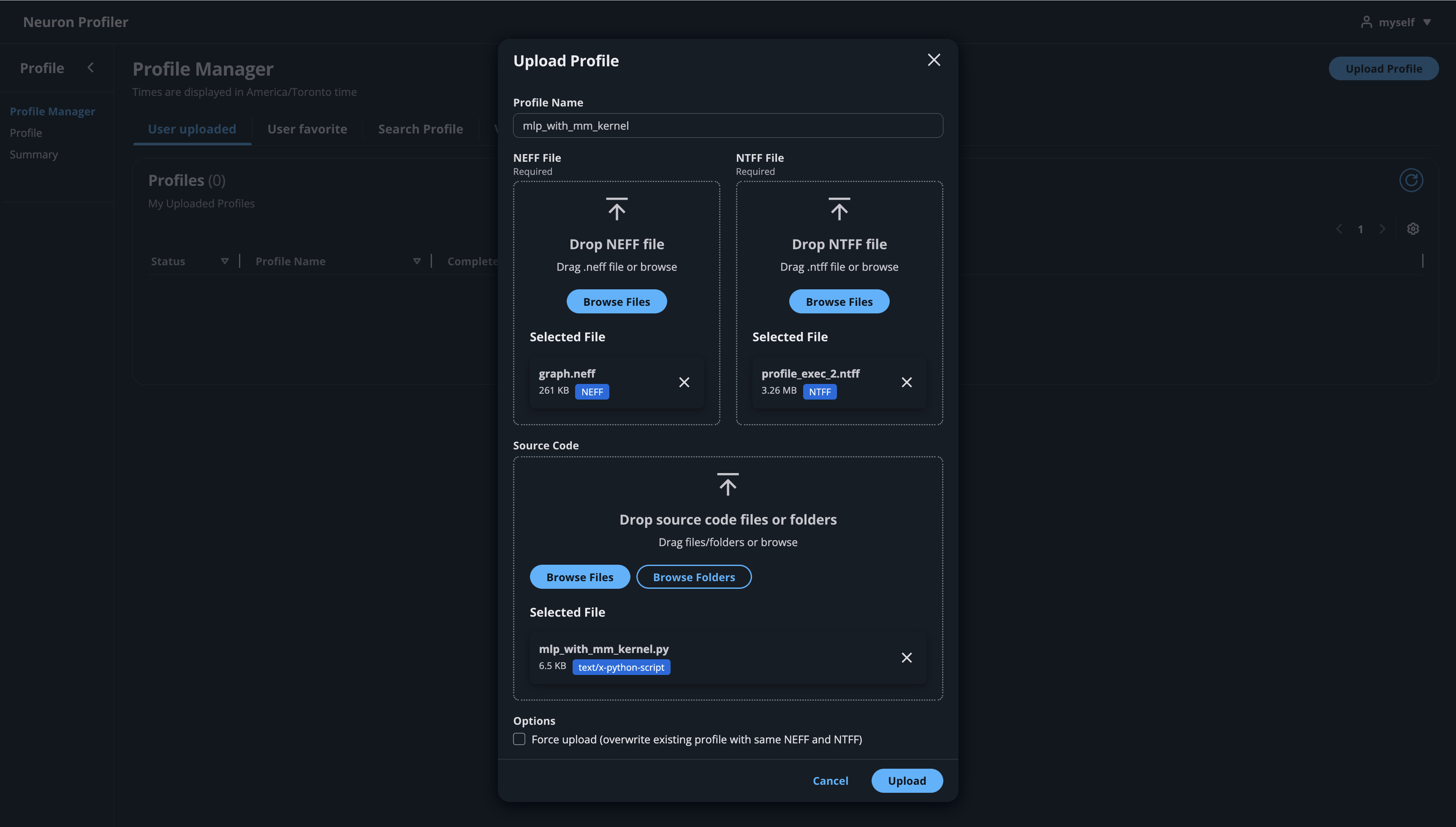
Click on the button “Upload Profile” to upload NEFF and NTFF files, and give a meaningful name to your profile. Selecting a source code folder for code linking is optional.
After the files are uploaded and processed, you will be able to open the profile from the list.
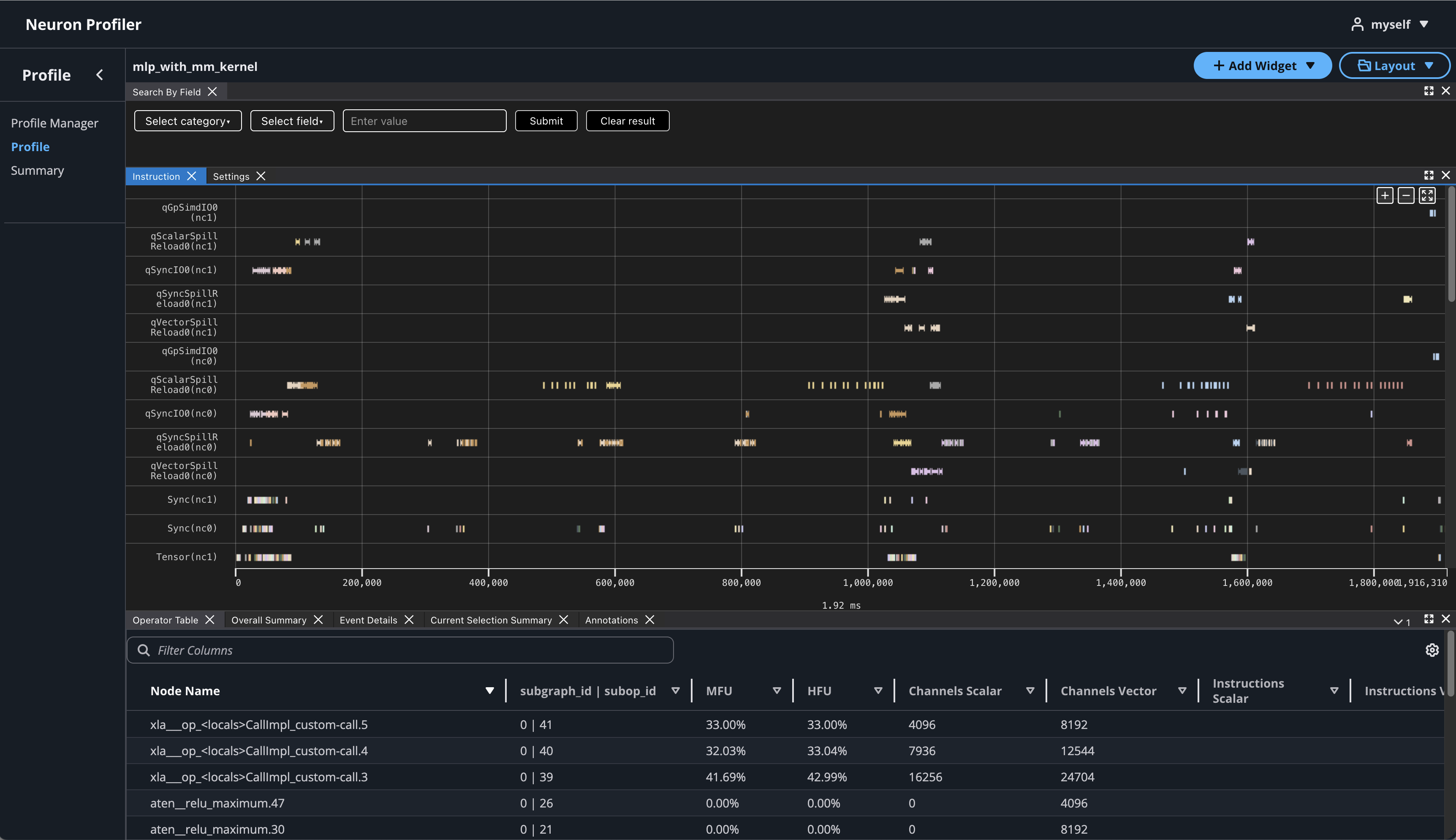
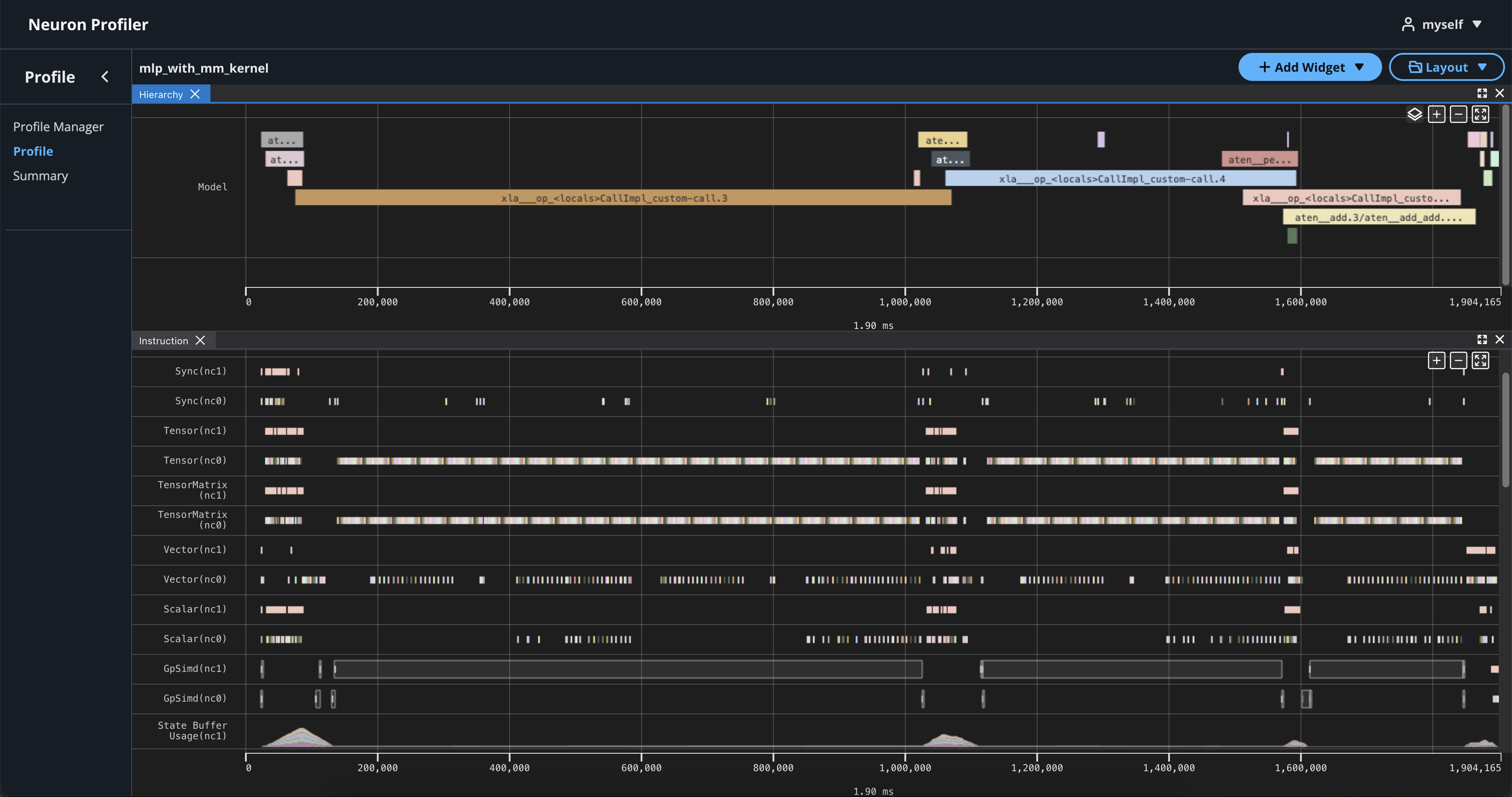
If you click on the name of your profile in Profile Name column, it will navigate to profile page
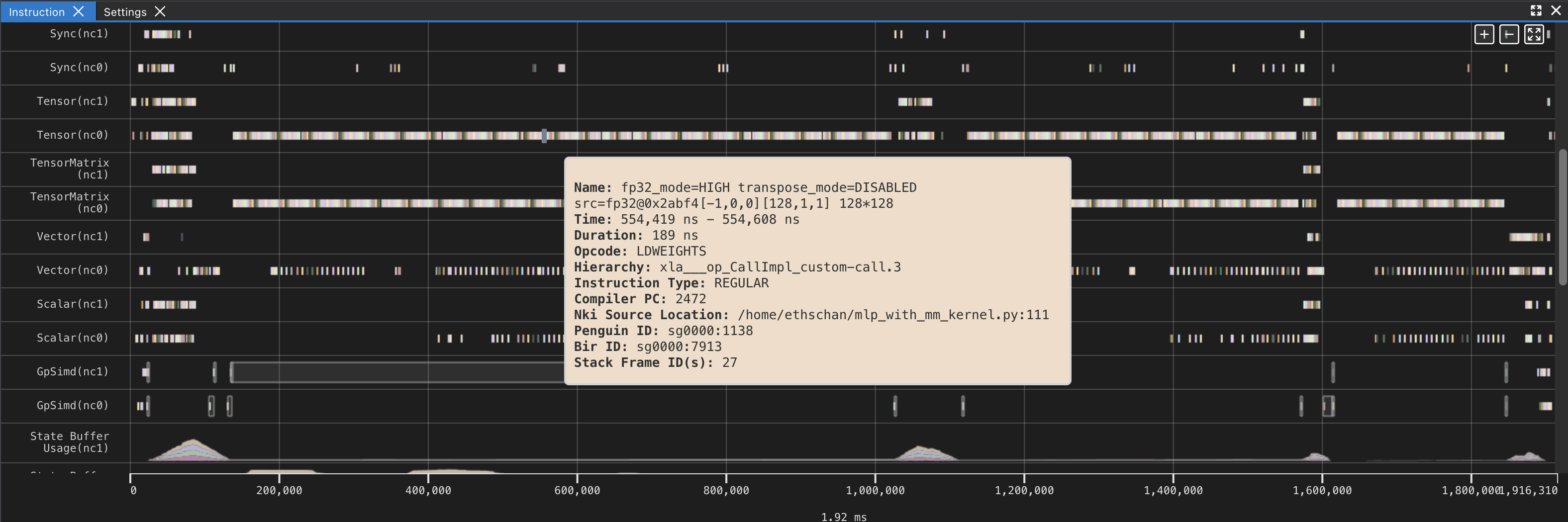
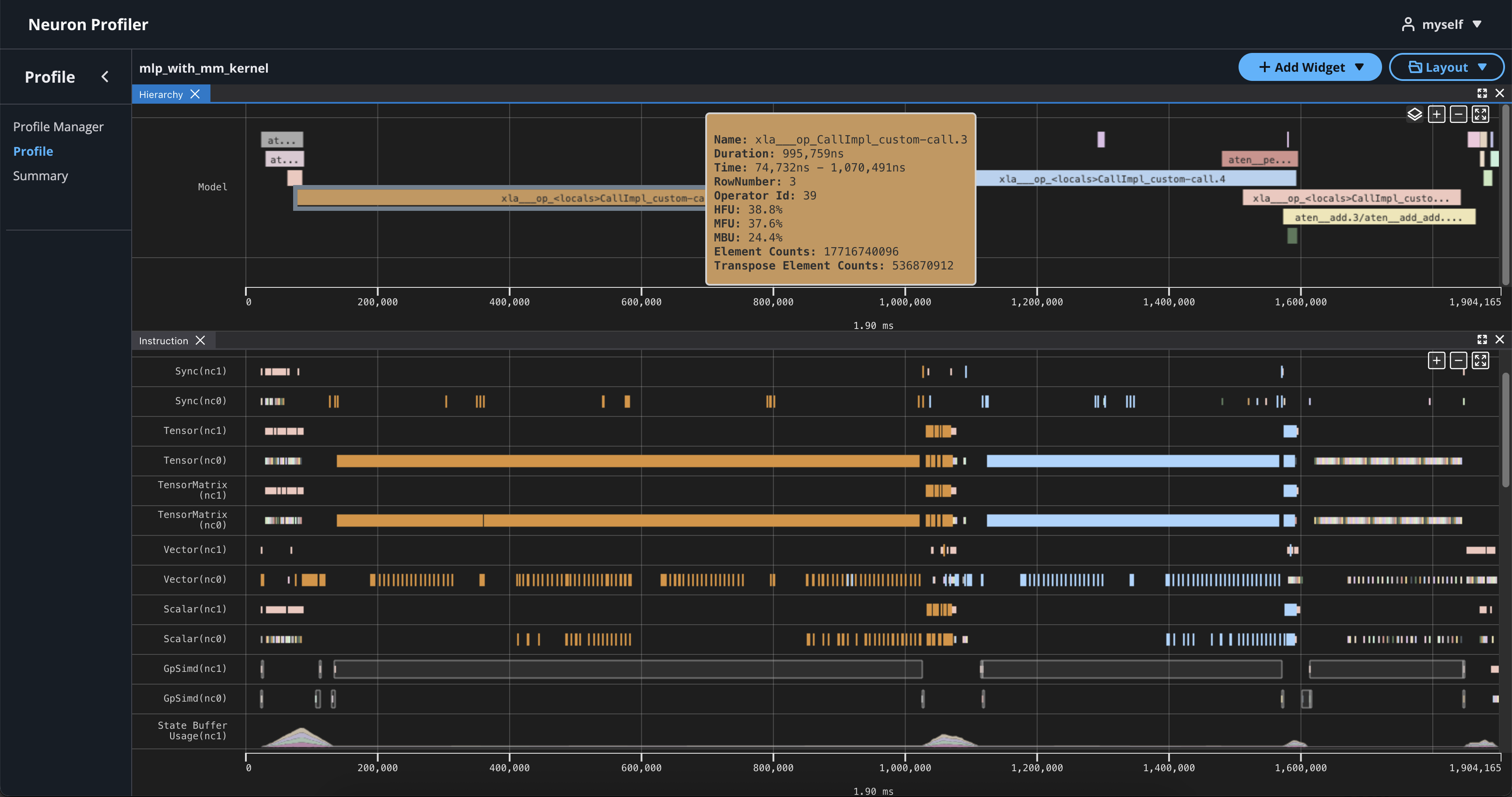
If you hover over any engine instruction in the timeline with your mouse, you will see instruction details in a pop-up box.
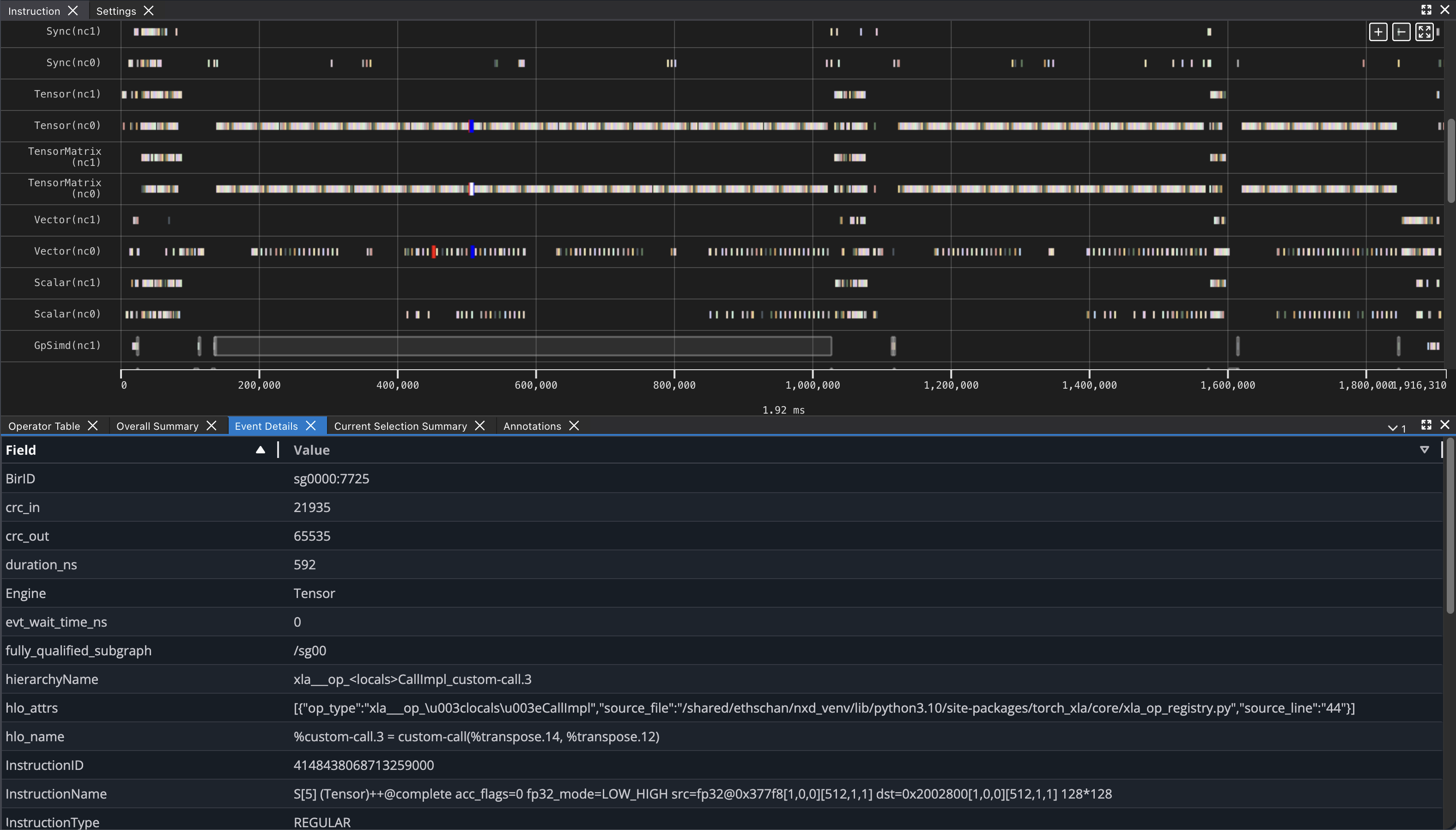
If you click on any engine instruction in the timeline with your mouse, you will see event details in a panel below the timeline.
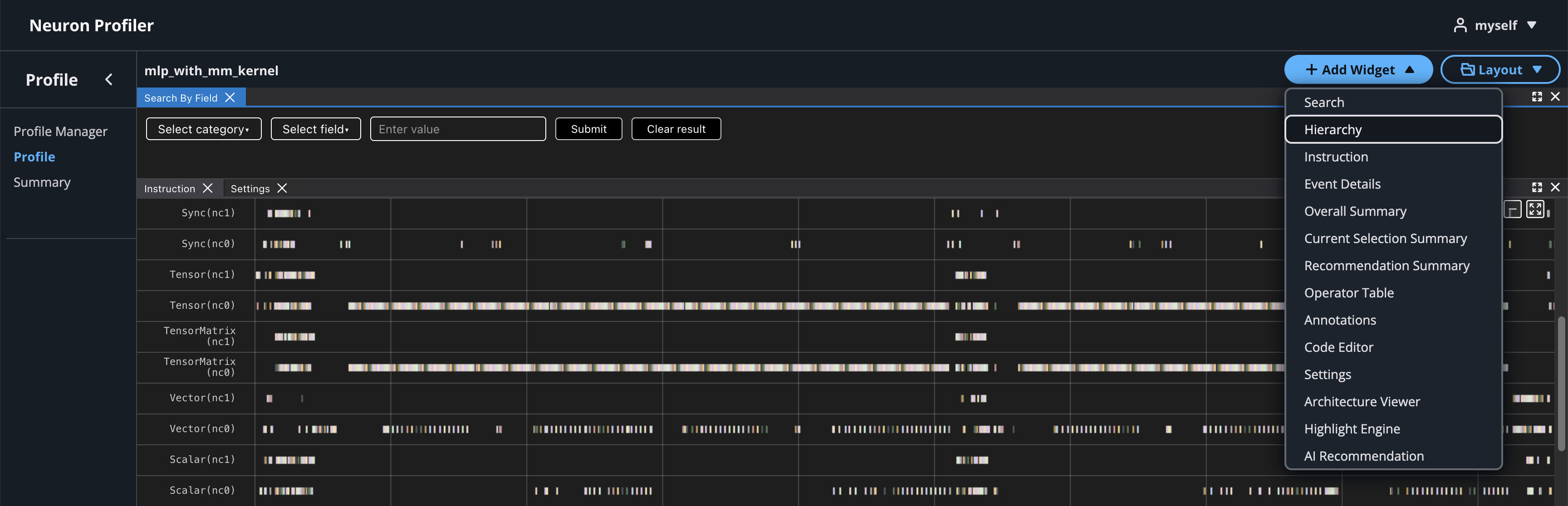
To view hierarchy of this profile, click on Add Widget and select Hierarchy. For more details, see the Hierarchy Viewer documentation.
Using the Profiler’s flexible layout support, you can drag and group every widget into any panel of your choice to customize the layout for your workflow.
View NKI Source Code in Neuron Profile#
You can optionally include your NKI source code files for display in Neuron Profile. When provided, Neuron Profile loads the source code into an integrated viewer, displayed side-by-side with the execution timeline in the web UI. This makes it easier to navigate between the instruction trace and the corresponding NKI source code, and to track the exact version of the code that generated the profile. For more details on source code linking, see the Source Code Viewer documentation.
Note
Even if you don’t upload the source code, the NKI source filename and line number remain available in the instruction detail view as noted in View Neuron Profile UI.
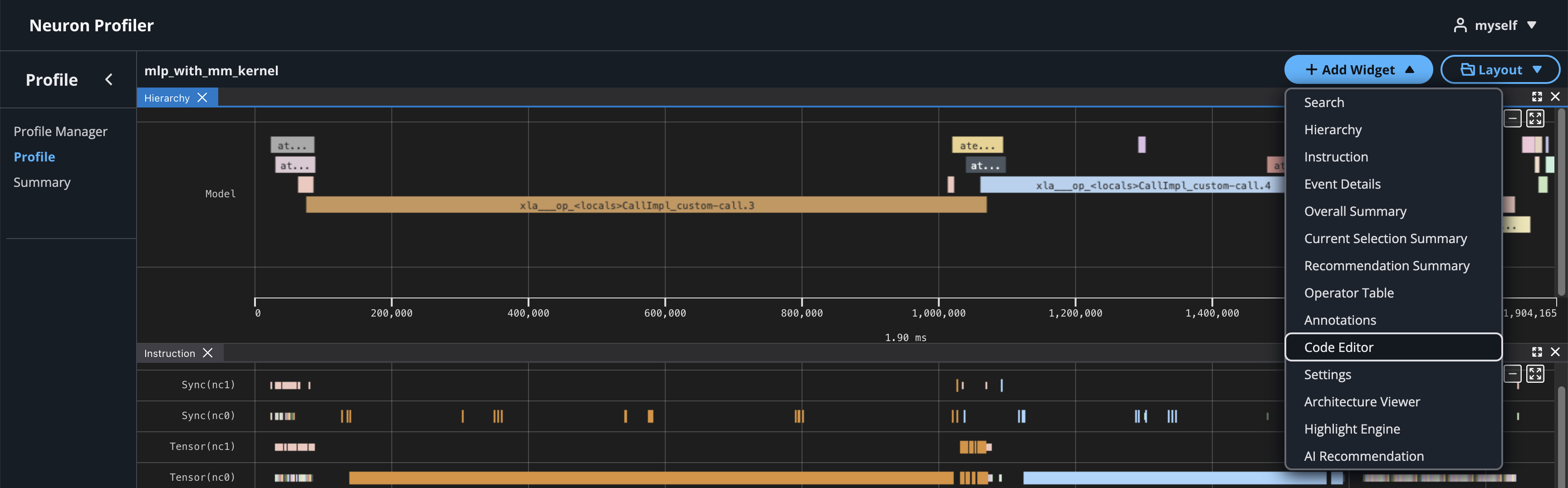
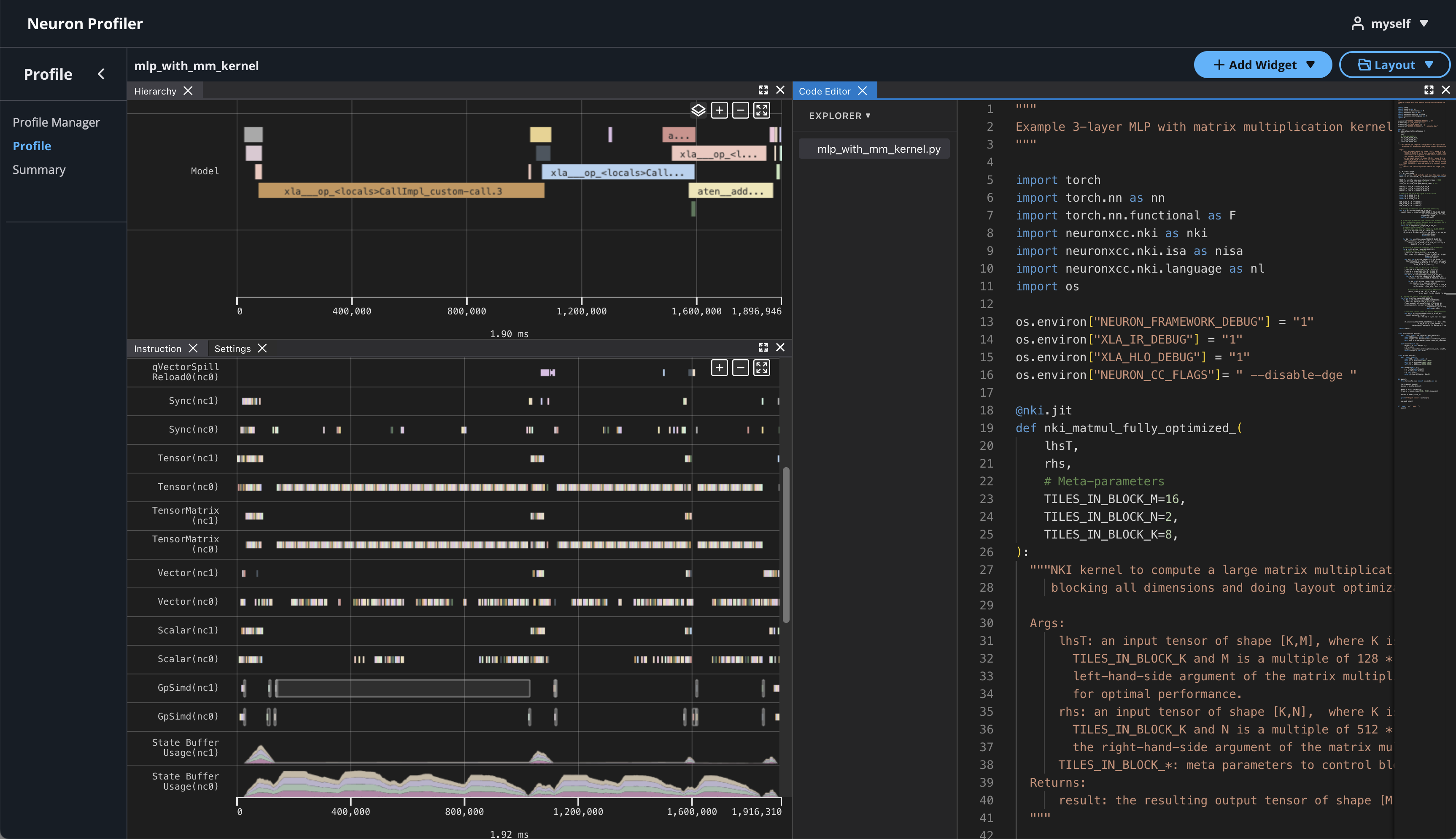
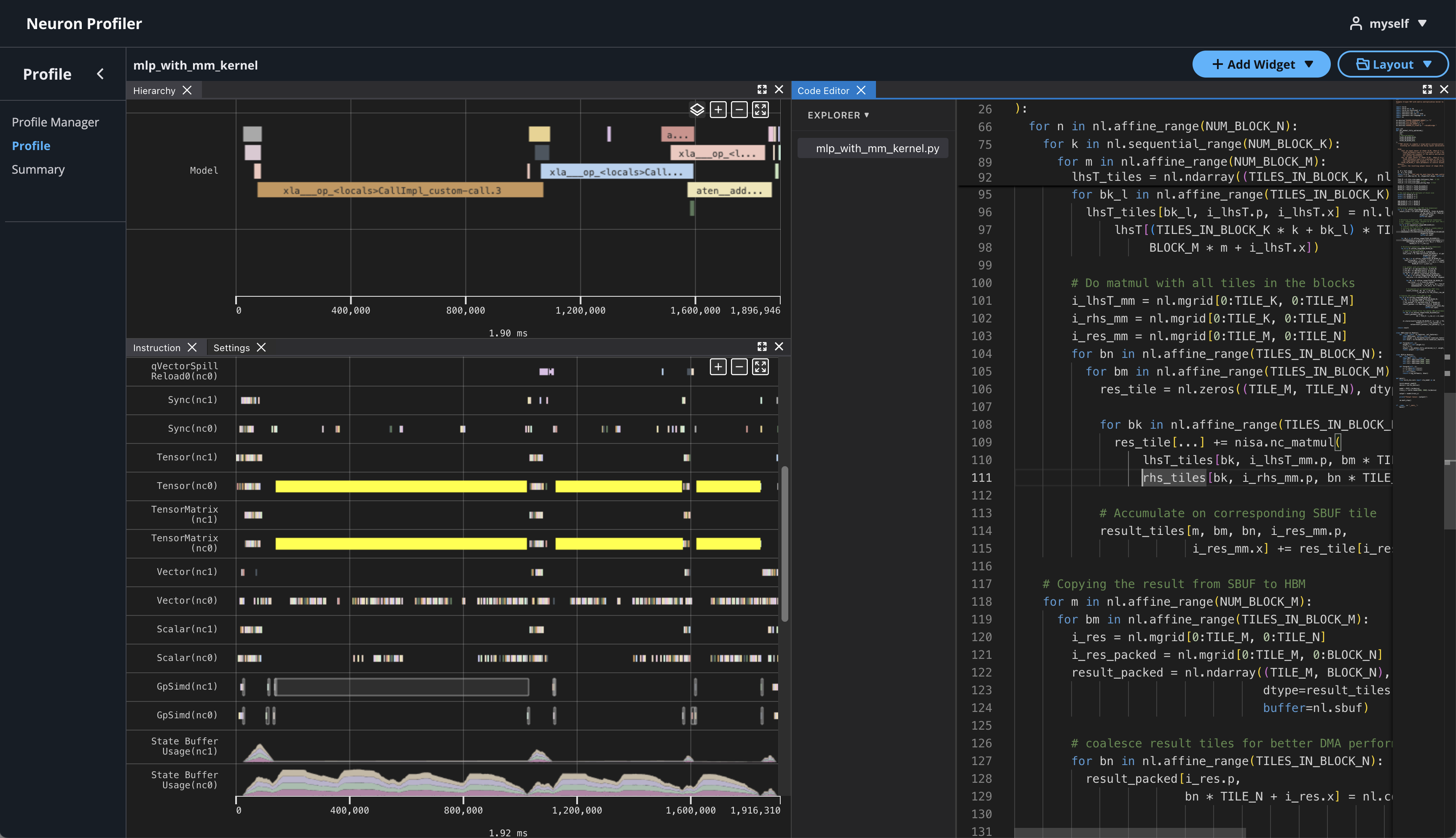
If source code is uploaded with NEFF and NTFF file, you will be able to see the source code in the code editor. To open the code editor, click on Add Widget and select Code Editor.
The code editor will be open on the right-hand side.
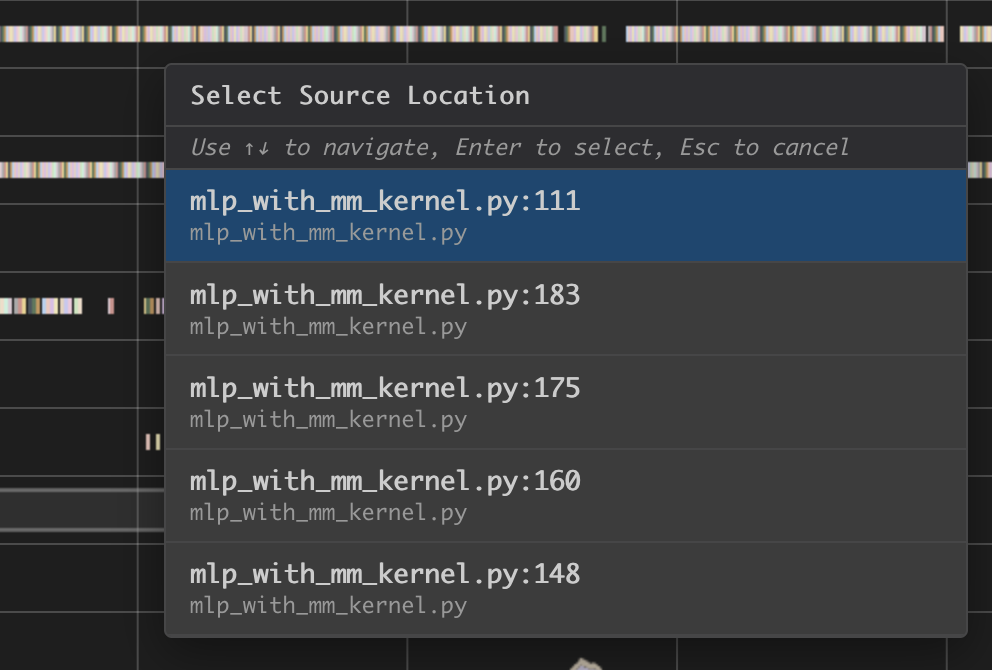
Hover on an instruction that has NKI source location and Command + left click on Mac (Ctrl + right click on Windows), and it will jump to the line of the source code and highlight all of instructions related to this line.
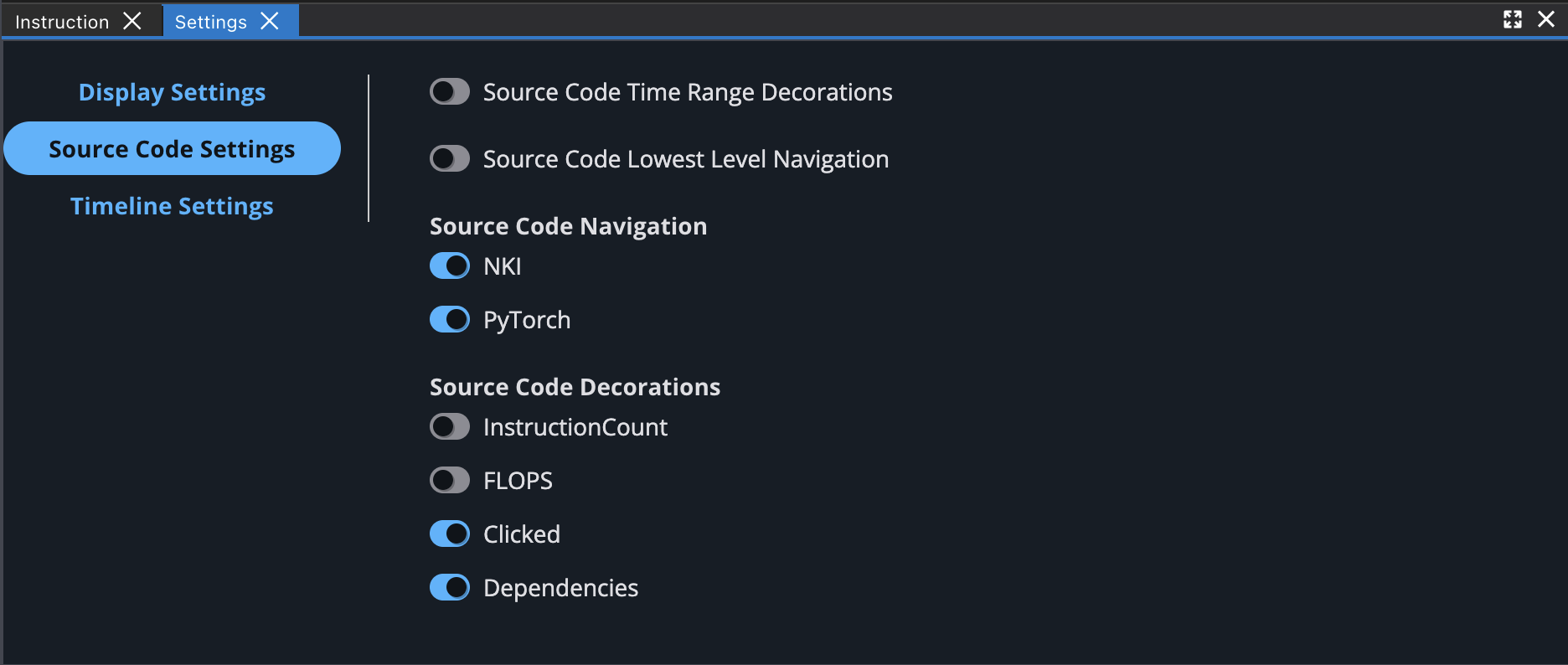
You can also enable different source code decorations in Source Code Settings.
Next Steps#
Great! Now that you’ve learned how to profile an NKI kernel, it’s time to take this further:
Dive into the NKI Performance Guide to discover techniques for making your kernels faster and more efficient.
Explore the NKI sample kernels to see real-world examples of high-performance kernel implementations — and get inspiration for your own NKI kernels.
Learn more about the Neuron Explorer viewers to deepen your profiling analysis:
Device Trace Viewer — Explore hardware-level execution with timeline view, operator table, and event details.
Hierarchy Viewer — Visualize execution from model layers down to hardware operations.
Source Code Viewer — Navigate between source code and profile data with bidirectional linking.
Summary Viewer — Get high-level performance insights and optimization recommendations.
AI Recommendation Viewer — Get AI-powered bottleneck analysis and optimization suggestions for NKI profiles.
By combining profiling insights with optimization strategies and practical examples, you’ll be well prepared to write NKI kernels that leverage Neuron hardware in an efficient way.
This document is relevant for: Trn2, Trn3