This document is relevant for: Trn2, Trn3
AveragePool2D#
In this tutorial, we examine a case of dimensionality reduction. We implement a 2D AveragePool operation, which is used in many vision neural networks. In doing so, we learn about:
NKI syntax and programming model.
multi-dimensional memory access patterns in NKI.
The 2D AveragePool operation takes
C x [H,W] matrices and reduces each matrix along the H and W
axes. To leverage free-dimension flexible indexing, we can map the C
(parallel) axis to the P dimension and H/W (contraction)
axes to the F dimension.
Performing such a 2D pooling operation requires a 4D memory access
pattern in the F dimension, with reduction along two axes.
Figure
below illustrates the input and output tensor layouts.
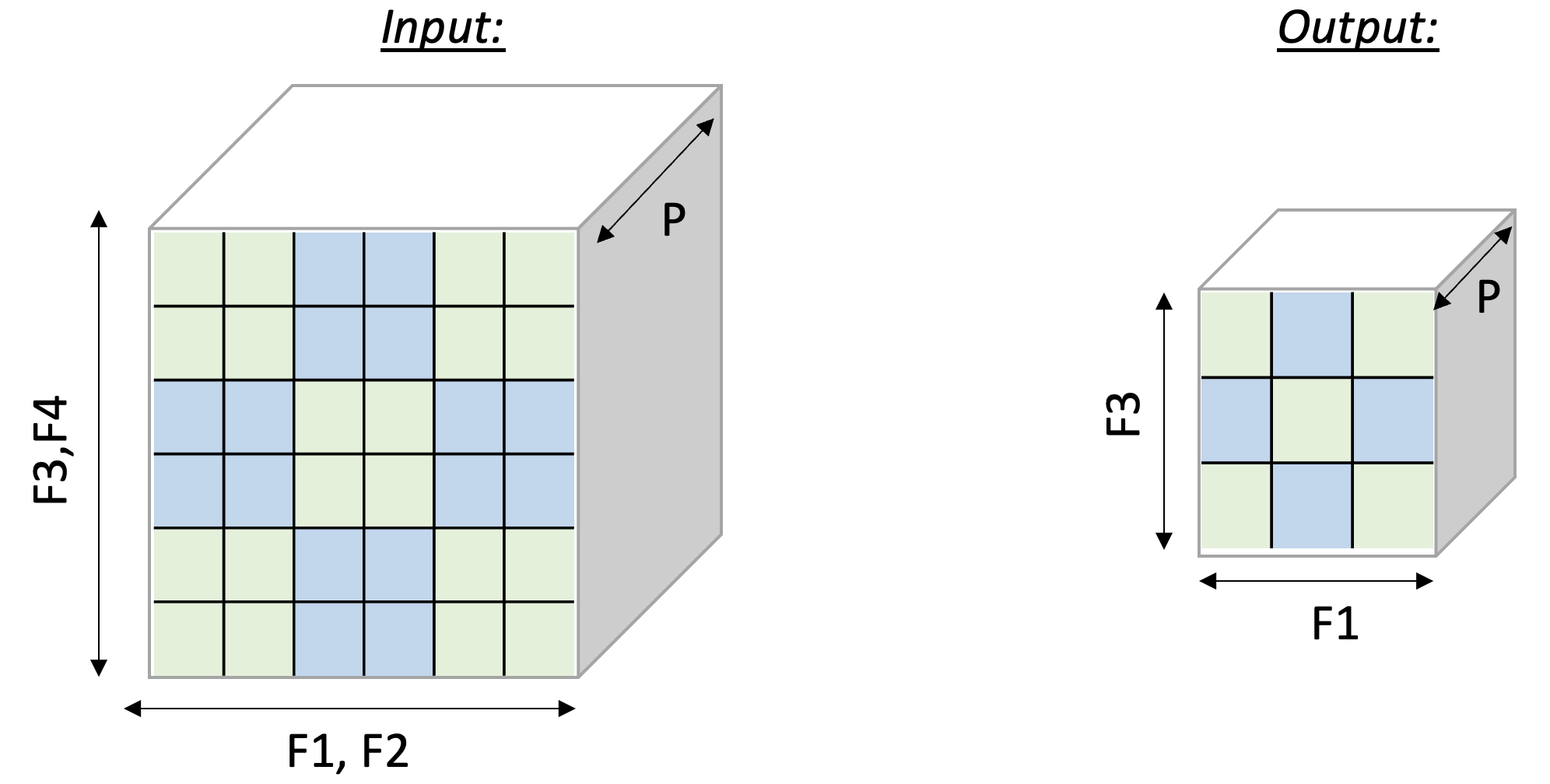
Fig. 23 2D-Pooling Operation (reducing on axes F2 and F4)#
PyTorch#
Compute kernel#
1import nki
2import nki.isa as nisa
3import nki.language as nl
4from nki.typing import tensor
5
6@nki.jit
7def tensor_avgpool_kernel(in_tensor, pool_size):
8 """NKI kernel to compute a 2D avg-pool operation
9
10 Args:
11 in_tensor: an input tensor, of shape C x H x W
12 pool_size: an integer representing a (square) pool-window size
13
14 Return:
15 out_tensor: the resulting output tensor, of shape C x (H/pool_size) x (W/pool_size)
16 """
17
18 # Get input/output dimensions
19 sz_cin, sz_hin, sz_win = in_tensor.shape
20 sz_hout = sz_hin // pool_size
21 sz_wout = sz_win // pool_size
22 # Create output tensor shared between all SPMD instances as result tensor
23 out_tensor = nl.ndarray((sz_cin, sz_hout, sz_wout), dtype=in_tensor.dtype,
24 buffer=nl.shared_hbm)
25
26 # Set relevant sizes
27 sz_p = sz_cin
28 sz_pool = pool_size
29
30 # Generate pool access pattern to create a 5D view:
31 # [sz_p, sz_hout, sz_wout, sz_pool, sz_pool]
32 # The pool dimensions are placed last so we can reduce over them.
33
34 # Load input data from external memory to on-chip memory
35 in_tile = nl.ndarray(in_tensor.shape, dtype=in_tensor.dtype, buffer=nl.sbuf)
36 nisa.dma_copy(dst=in_tile, src=in_tensor)
37
38 # Perform the pooling operation using an access pattern view:
39 # The .ap() creates a strided 5D view of the 3D input tile,
40 # grouping elements into pool windows for reduction.
41 pool_view = in_tile.ap([
42 [sz_hin * sz_win, sz_p], # partition stride
43 [sz_pool * sz_win, sz_hin // sz_pool], # outer row stride
44 [sz_pool, sz_win // sz_pool], # outer col stride
45 [sz_win, sz_pool], # inner row stride (within pool window)
46 [1, sz_pool], # inner col stride (within pool window)
47 ])
48 sum_tile = nl.sum(pool_view, axis=[3, 4])
49 out_tile = nl.ndarray(sum_tile.shape, dtype=sum_tile.dtype, buffer=nl.sbuf)
50 nisa.tensor_scalar(dst=out_tile, data=sum_tile, op0=nl.multiply,
51 operand0=1.0 / (pool_size * pool_size))
52
53 # Store the results back to hbm
54 nisa.dma_copy(dst=out_tensor, src=out_tile)
55
56 # Transfer the ownership of `out_tensor` to the caller
57 return out_tensor
Launching kernel and testing correctness#
To execute the kernel, we prepare tensors in_tensor and call tensor_avgpool_kernel:
1import torch
2import torch_xla
3
4if __name__ == "__main__":
5 device = torch_xla.device()
6
7 # Now let's run the kernel
8 POOL_SIZE = 2
9 C, HIN, WIN = 2, 6, 6
10 HOUT, WOUT = HIN//POOL_SIZE, WIN//POOL_SIZE
11
12 in_tensor = torch.arange(C * HIN * WIN, dtype=torch.bfloat16).reshape(C, HIN, WIN).to(device=device)
13 out_nki = torch.zeros((C, HOUT, WOUT), dtype=torch.bfloat16).to(device=device)
14
15 out_nki = tensor_avgpool_kernel(in_tensor, POOL_SIZE)
16
17 out_torch = torch.nn.functional.avg_pool2d(in_tensor, POOL_SIZE, POOL_SIZE)
18
19 print(in_tensor, out_nki, out_torch) # an implicit XLA barrier/mark-step
20
21 if (out_nki == out_torch).all():
22 print("NKI and Torch match")
23 else:
24 print("NKI and Torch differ")
JAX#
Compute kernel#
Let’s reuse the same NKI kernel implementation defined for PyTorch above:
1import nki
2import nki.isa as nisa
3import nki.language as nl
4from nki.typing import tensor
5
6@nki.jit
7def tensor_avgpool_kernel(in_tensor, pool_size):
8 """NKI kernel to compute a 2D avg-pool operation
9
10 Args:
11 in_tensor: an input tensor, of shape C x H x W
12 pool_size: an integer representing a (square) pool-window size
13
14 Return:
15 out_tensor: the resulting output tensor, of shape C x (H/pool_size) x (W/pool_size)
16 """
17
18 # Get input/output dimensions
19 sz_cin, sz_hin, sz_win = in_tensor.shape
20 sz_hout = sz_hin // pool_size
21 sz_wout = sz_win // pool_size
22 # Create output tensor shared between all SPMD instances as result tensor
23 out_tensor = nl.ndarray((sz_cin, sz_hout, sz_wout), dtype=in_tensor.dtype,
24 buffer=nl.shared_hbm)
25
26 # Set relevant sizes
27 sz_p = sz_cin
28 sz_pool = pool_size
29
30 # Generate pool access pattern to create a 5D view:
31 # [sz_p, sz_hout, sz_wout, sz_pool, sz_pool]
32 # The pool dimensions are placed last so we can reduce over them.
33
34 # Load input data from external memory to on-chip memory
35 in_tile = nl.ndarray(in_tensor.shape, dtype=in_tensor.dtype, buffer=nl.sbuf)
36 nisa.dma_copy(dst=in_tile, src=in_tensor)
37
38 # Perform the pooling operation using an access pattern view:
39 # The .ap() creates a strided 5D view of the 3D input tile,
40 # grouping elements into pool windows for reduction.
41 pool_view = in_tile.ap([
42 [sz_hin * sz_win, sz_p], # partition stride
43 [sz_pool * sz_win, sz_hin // sz_pool], # outer row stride
44 [sz_pool, sz_win // sz_pool], # outer col stride
45 [sz_win, sz_pool], # inner row stride (within pool window)
46 [1, sz_pool], # inner col stride (within pool window)
47 ])
48 sum_tile = nl.sum(pool_view, axis=[3, 4])
49 out_tile = nl.ndarray(sum_tile.shape, dtype=sum_tile.dtype, buffer=nl.sbuf)
50 nisa.tensor_scalar(dst=out_tile, data=sum_tile, op0=nl.multiply,
51 operand0=1.0 / (pool_size * pool_size))
52
53 # Store the results back to hbm
54 nisa.dma_copy(dst=out_tensor, src=out_tile)
55
56 # Transfer the ownership of `out_tensor` to the caller
57 return out_tensor
In order to pass pool_size as a compile time constant, we pass pool_size as kwargs.
out_nki = tensor_avgpool_kernel(in_array, pool_size=POOL_SIZE)
We write a reference JAX implementation of AveragePool2D as JAX does
not have a primitive for it.
1import jax.numpy as jnp
2
3# Reference JAX implementation
4def jax_average_pool_2D(in_tensor, pool_size):
5 c, h_in, w_in = in_tensor.shape
6 reshaped = in_tensor.reshape(c, h_in // pool_size, pool_size, w_in // pool_size, pool_size)
7 return jnp.nanmean(reshaped, axis=(2, 4))
Launching kernel and testing correctness#
To execute the kernel, we prepare array in_array and invoke the kernel caller function tensor_avgpool_kernel:
1if __name__ == "__main__":
2 POOL_SIZE = 2
3 C, HIN, WIN = 2, 6, 6
4 HOUT, WOUT = HIN//POOL_SIZE, WIN//POOL_SIZE
5
6 in_array = jnp.arange(C * HIN * WIN, dtype=jnp.float32).reshape(C, HIN, WIN)
7
8 out_nki = tensor_avgpool_kernel(in_array, pool_size=POOL_SIZE)
9 out_jax = jax_average_pool_2D(in_array, pool_size=POOL_SIZE)
10
11 print(in_array, out_nki, out_jax)
12
13 if jnp.allclose(out_nki, out_jax):
14 print("NKI and JAX match")
15 else:
16 print("NKI and JAX differ")
Download All Source Code#
Click the links to download source code of the kernels and the testing code discussed in this tutorial.
NKI baremetal implementation:
average_pool2d_nki_kernels.py- PyTorch implementation:
average_pool2d_torch.py You must also download
average_pool2d_nki_kernels.pyinto the same folder to run this PyTorch script.
- PyTorch implementation:
- JAX implementation:
average_pool2d_jax.py You must also download
average_pool2d_nki_kernels.pyinto the same folder to run this JAX script.
- JAX implementation:
You can also view the source code in the GitHub repository nki_samples
Example usage of the scripts:#
Run NKI baremetal implementation:
python3 average_pool2d_nki_kernels.py
Run PyTorch implementation:
python3 average_pool2d_torch.py
Run JAX implementation:
python3 average_pool2d_jax.py
This document is relevant for: Trn2, Trn3